Chromatin & DNA Modeling
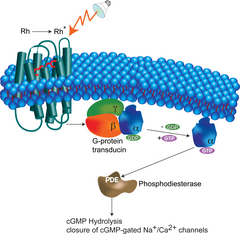
Cytoskeleton Dynamics and Cell Motility

| Project Title | Short Summary | Lab Members | |||
|
|
|
|
|
||

|
Chromatin & DNA Modeling |
DNA associates with histon proteins in higher eukaryotic cells, compactifying million-fold. The resulting chromatin fibers are very dynamic, regulating gene transcription. Using all-atom and coarse-grained molecular dynamics simulations, we are developing new insights into the chromatin fiber structure and stability. Learn more... | Haiqing Zhao, Hao Wu, Mary Pitman | ||
|
|
|
|
|
||

|
Cytoskeleton Dynamics and Cell Motility |
Employing tools from non-equilibrium statistical physics, we are building models of signal transduction networks. In particular, we are interested in spatio-temporal regulation of eukaryotic cell motility. We have developed models for actin based cellular organelles, such as filopodia -- the fingerlike sensorial membrane protrusions, -- and lamellipodia -- larger flat actin-based cell motility structures Learn more... | James Komianos, Aravind Chandrasekaran, Qin Ni, Haoran Ni | ||
|
|
|
|
|
||

|
Native Protein Dynamics |
We are developing an integrated system of multi-scale computational techniques, from very coarse-grained to all-atom models, to describe the native dynamics of various enzymes and signaling proteins. In a related project, we are working on new free energy computational methods for proteins. Learn more... | Haiqing Zhao, Hao Wu | ||
|
|
|
|
|
Last updated: January 23rd, 2017